|
#2
30th September 2017, 11:54 AM
| |||
| |||
| Re: RCSB Ligand Explorer Viewer
As you want to know details of State University of New Jersey, FAQs on Ligand Explorer software of RCSB PDB, here I will get information for you. Here are some FAQs How to rotate or move the Structures? To quickly get information on structural details you may mouse over structural features in the molecular viewer window as well as the sequence viewer at the top. Information regarding structural details such as residue number and element type is displayed in the status bar at the bottom of the viewer. How display Interactions with the Ligand? First, select a ligand from the left hand menu. If you launched Ligand Explorer from the Ligand Chemical Component widget, a ligand is already selected. Second, click the check boxes next to the list of interactions to turn on the interactions. What types of Interactions are available? Hydrogen bonds (pink lines) Hydrophobic interactions (green lines) Bridged hydrogen bonds (water mediated hydrogen bonds) (blue lines) Metal interactions (gray lines) Neighbor residues (interactions with surrounding residues) FAQs on Ligand Explorer software   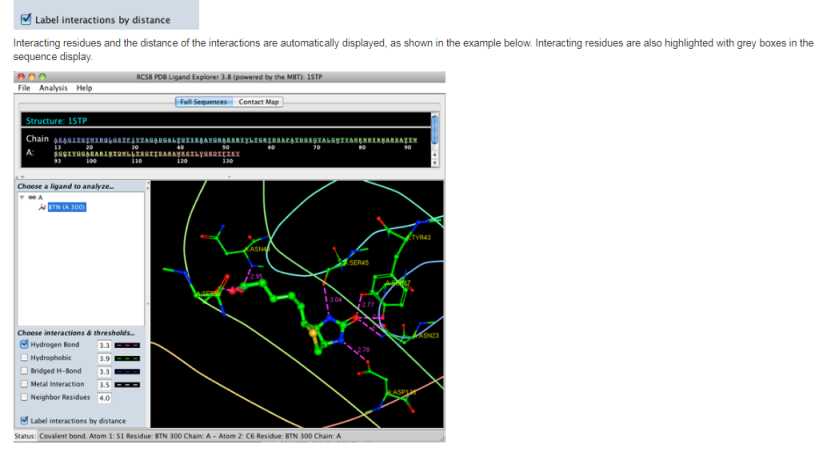  Address;- State University of New Jersey Center for Integrative Proteomics Research 174 Frelinghuysen Rd Piscataway, NJ 08854-8076 |